Thesis - Neural Processing of 3-D VisionMy current project investigates how distinct brain regions are connected, providing insight into how complex neural networks are formed. In this work, we use magnetic resonance imaging (MRI) and diffusion weighted imaging (DWI) to infer connections between different brain regions through probabilistic inference. Using novel metrics that we are developing, we are able to infer whether or not brain regions are connected, how strongly they are connected, and construct probabilistic flow diagrams of brain networks.
Currently, I am using this method to investigate the cortical network within the dorsal visual stream (i.e., the ‘where’ pathway), which is responsible for transforming two-dimensional retinal images into three-dimensional (3D) visual perception. In the future we plan to extend this method to elucidate other cortical pathways, such as the ventral visual stream (i.e., the ‘what’ pathway) and the pathways that bridge 3D vision and motor outputs (e.g., reaching and grasping). |
Real-Time Experimental Control with Graphical User Interface (REC-GUI)
The REC-GUI Framework -
Modern neuroscience research often requires the coordination of multiple processes such as stimulus generation, real-time experimental control, as well as behavioral and neural measurements. The technical demands required to simultaneously manage these processes with high temporal fidelity limits the number of labs capable of performing such work. The Real-Time Experimental Control with Graphical User Interface (REC-GUI) Framework is an open-source, network-based parallel processing framework that lowers this barrier. The Framework offers multiple advantages, including: (i) a modular design that is agnostic to coding language(s) and operating system(s) to maximize experimental flexibility and minimize researcher effort, (ii) simple interfacing to connect multiple measurement and recording devices, (iii) high temporal fidelity by dividing task demands across CPUs, and (iv) real-time control using a fully customizable and intuitive GUI.
Modern neuroscience research often requires the coordination of multiple processes such as stimulus generation, real-time experimental control, as well as behavioral and neural measurements. The technical demands required to simultaneously manage these processes with high temporal fidelity limits the number of labs capable of performing such work. The Real-Time Experimental Control with Graphical User Interface (REC-GUI) Framework is an open-source, network-based parallel processing framework that lowers this barrier. The Framework offers multiple advantages, including: (i) a modular design that is agnostic to coding language(s) and operating system(s) to maximize experimental flexibility and minimize researcher effort, (ii) simple interfacing to connect multiple measurement and recording devices, (iii) high temporal fidelity by dividing task demands across CPUs, and (iv) real-time control using a fully customizable and intuitive GUI.
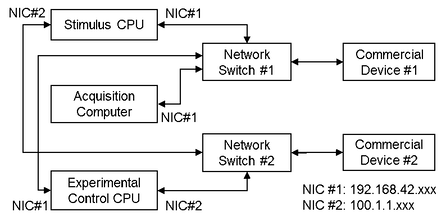
Experimental systems often require multiple pieces of specialized hardware produced by different companies (e.g., commercial devices 1 and 2). Network-based communications with such hardware must often occur over non-configurable, predefined subgroups of IP addresses. In the figure above, commercial devices 1 and 2 have non-configurable, predefined subgroups of IP addresses that cannot be routed over the same network interface card (NIC) without additional configuration of the routing tables. In such cases, a simple way to set up communication with the hardware is to use multiple parallel networks with a dedicated network switch for each piece of hardware. For this hypothetical configuration, this requires two network switches, and both the stimulus and experimental control CPUs require two NICs assigned to different subgroups of IP addresses. With this configuration, the stimulus and experimental control CPUs can both directly communicate with commercial devices 1 and 2.
Machine Learning - Support Vector Machine
In our visual system, there are two main pathways; (i) the 'what' pathway and (ii) the 'where' pathway. Neurons in the 'what' pathway encode for specific objects such as a chairs and faces, whereas neurons in the 'where' pathway encode for the object's location and orientation in three-dimensional (3D) space.
Our lab studies the information encoded by different brain areas within the 'where' pathway by recording from multiple neurons simultaneously. We present objects at different orientations and record how each neuron responds to the stimuli. To quantify the response of each neuron, we calculate the average firing rate, or the number of times the neuron fired an action potential while the stimulus was present divided by the time the stimulus was present (i.e., firing rate: spikes per second). From this data we can use a machine learning model to test the hypothesis that these neurons in the 'where' pathway actually encode for the orientation of the object in space.
Our lab studies the information encoded by different brain areas within the 'where' pathway by recording from multiple neurons simultaneously. We present objects at different orientations and record how each neuron responds to the stimuli. To quantify the response of each neuron, we calculate the average firing rate, or the number of times the neuron fired an action potential while the stimulus was present divided by the time the stimulus was present (i.e., firing rate: spikes per second). From this data we can use a machine learning model to test the hypothesis that these neurons in the 'where' pathway actually encode for the orientation of the object in space.
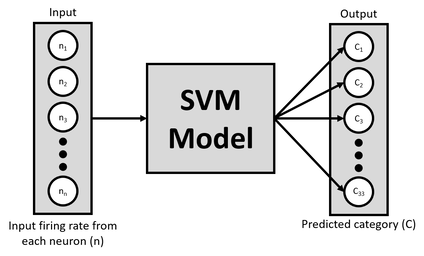
Here, we use a support vector machine (SVM; above figure) to classify the neuron's response to each stimuli. Each neuron is a different feature in the model, and all neurons were shown the same type of stimuli. In our experimental design, we present 33 different orientations and repeat this multiple times to have many repetitions of each stimulus. The goal of the model is to classify which stimulus was presented given some firing rate for each of the neurons. If the model can successfully learn this task, this would suggest that the specific brain areas within the 'where' pathway are indeed encoding an object's orientation.
Computing Brain Connectivity
In this work, we use magnetic resonance imaging (MRI) and diffusion weighted imaging (DWI) to infer connections between different brain regions through probabilistic inference. Using novel metrics that we are developing, we are able to infer whether or not brain regions are connected, how strongly they are connected, and construct probabilistic flow diagrams of brain networks. We then use graph theory to analyze information flow throughout subnetworks within the brain.
Currently, I am using this method to investigate the cortical network within the dorsal visual stream (i.e., the ‘where’ pathway), which is responsible for transforming two-dimensional retinal images into three-dimensional (3D) visual perception. In the future we plan to extend this method to elucidate other cortical pathways, such as the ventral visual stream (i.e., the ‘what’ pathway, similar to object recognition) and the pathways that bridge 3D vision and motor outputs (e.g., reaching and grasping).
Currently, I am using this method to investigate the cortical network within the dorsal visual stream (i.e., the ‘where’ pathway), which is responsible for transforming two-dimensional retinal images into three-dimensional (3D) visual perception. In the future we plan to extend this method to elucidate other cortical pathways, such as the ventral visual stream (i.e., the ‘what’ pathway, similar to object recognition) and the pathways that bridge 3D vision and motor outputs (e.g., reaching and grasping).
Get In Touch
If you have any questions about my work, please feel free to contact me.
I would be happy to answer any questions that you may have.
I would be happy to answer any questions that you may have.